TSK and Probit Models
Source:vignettes/articles/TSK-and-Probit-Models.Rmd
TSK-and-Probit-Models.Rmd
library(drcHelper)
#> Loading required package: drc
#> Loading required package: MASS
#> Loading required package: drcData
#>
#> 'drc' has been loaded.
#> Please cite R and 'drc' if used for a publication,
#> for references type 'citation()' and 'citation('drc')'.
#>
#> Attaching package: 'drc'
#> The following objects are masked from 'package:stats':
#>
#> gaussian, getInitialTrimmed Spearman-Karber (TSK) Method
The Trimmed Spearman-Karber (TSK) method is a statistical technique used primarily in toxicology and ecotoxicology to estimate the median effective dose/concentration/rate (ED50, EC50, ER50) or median lethal concentration (LC50) from dose-response data. It builds on the original Spearman-Karber method, which is a non-parametric approach for calculating the point where 50% of the test subjects respond (e.g., dead or symptom present). The “trimmed” aspect, introduced by Hamilton et al. in 1977, enhances robustness by excluding extreme data points where the response proportions are near 0% or 100%, reducing bias from incomplete dose ranges.
How the TSK Approach Works
The TSK method approximates the mean of the underlying tolerance
distribution, assuming it is symmetric on a logarithmic scale (common
for concentrations). It processes data through several steps to compute
the ED50 (referred to as mu in the provided code), its
variance, geometric standard deviation (GSD), and confidence intervals.
Here’s a breakdown based on the algorithm in the code and standard
descriptions:
-
Input Validation and Data Preparation:
- Inputs include:
x(vector of doses or concentrations, must be positive and increasing),r(number of responses, e.g., deaths),n(number of subjects tested per dose),A(trim level, 0 ≤ A < 0.5, default 0), andconf(confidence level, 0.5 < conf < 1, default 0.95). - Doses are log-transformed (base 10) for symmetry. If the first dose is 0 (control), it’s handled by scaling responses or issuing warnings if non-zero.
- Proportions
p = r/nare calculated.
- Inputs include:
Below is an example data set from the R package ecotoxicology.
# Example data
data <- data.frame(
x = c(15.54, 20.47, 27.92, 35.98, 55.52),
r = c(0, 0, 0, 1, 20),
n = c(20, 20, 20, 19, 20)
)-
Smoothing:
- If proportions are not monotonically increasing (which can happen due to sampling variability), adjacent non-increasing values are averaged to enforce monotonicity. This prevents artifacts in estimation.
-
Scaling and Trimming:
- Proportions are scaled:
p_scaled = (p - A) / (1 - 2*A)to focus on the central response range (A to 1-A). - Data points where p ≤ A or p ≥ 1-A are trimmed. Linear interpolation adds virtual points at exactly 0% and 100% response if needed, ensuring the trimmed dataset spans from 0 to 1.
- This trimming avoids over-reliance on tails where variability is high or data is sparse.
- Proportions are scaled:
-
Estimation of the Median (mu):
- The trimmed log-doses and differences in proportions
(
delp) are used to compute the midpoint averages. - The log-median is
logmu = sum((x_i + x_{i+1})/2 * delp_i), a weighted average approximating the mean of the log-tolerance distribution. - The median dose is
mu = 10^logmu.
- The trimmed log-doses and differences in proportions
(
-
Variance and Confidence Intervals:
- Variance is calculated using specialized functions (V1 to V6 in the code) based on the positions of lower (L) and upper (U) bounds around the trim.
- These account for binomial variability in proportions and dose
spacings, scaled by
(2 - 4*A)^2. - GSD =
10^sqrt(Var), representing dispersion. - Confidence limits use the inverse error function: left =
mu * gsd^(-v), right =mu * gsd^v, wherev = sqrt(2) * erfinv(conf)for approximate normality on log scale.
-
Output and Visualization:
- Returns
mu,gsd,left, andright. - The code includes plotting: response curve, median point, and error bars for confidence.
- Returns
This method is computationally simple, non-iterative, and excels in handling datasets where full 0-100% response isn’t achieved.
# Load libraries for probit analyses
library(MASS)
library(drc)
# Compute TSK
tsk_result_ecotox <- ecotoxicology::TSK(data$x, data$r, data$n, A = 0, conf = 0.95)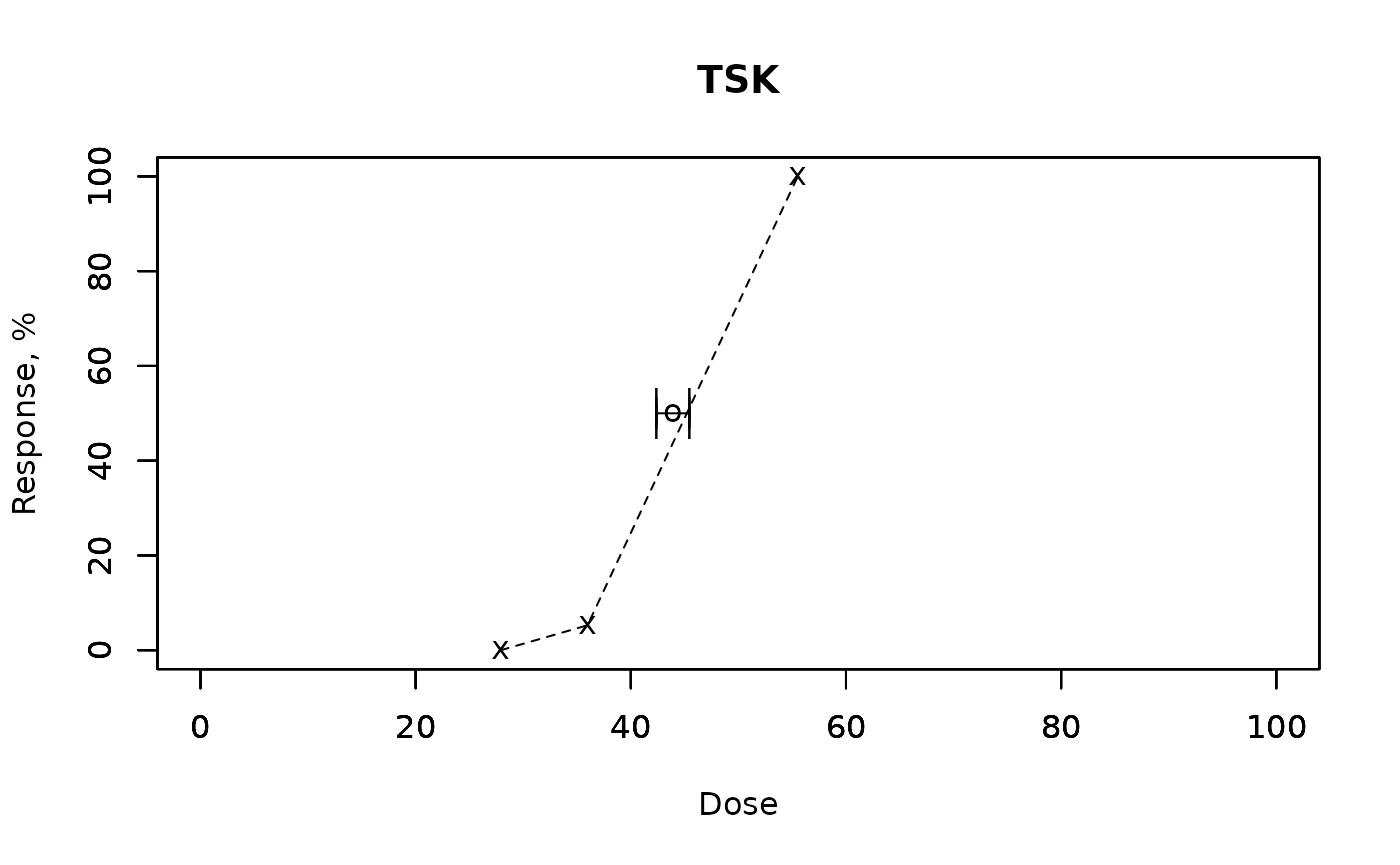
Validation against ecotoxicology::TSK
This part validate that the tsk function included in the drcHelper function computes TSK estimate and confidence intervals matching ecotoxicology::TSK.
library(drcHelper)
tsk_result_drc <- drcHelper::tsk(
x = data$x,
r = data$r,
n = data$n,
control = 0,
trim = 0,
conf.level = 0.95,
use.log.doses = TRUE
)
print("TSK Results from drcHelper:")
#> [1] "TSK Results from drcHelper:"
print(tsk_result_drc)
#>
#> Trimmed Spearman-Karber method using 0 percent trim
#>
#> Data was not smoothed
#> Calculation done using the logs of the doses
#> Estimated LD50: 43.89339 GSD of estimate: 1.017763
#> 95 percent confidence interval on LD50:
#> 42.40451 45.43455
comparison_table <- data.frame(
Metric = c("LD50/Mu", "GSD", "CI Lower", "CI Upper"),
ecotoxicology_TSK = c(tsk_result_ecotox$mu, tsk_result_ecotox$gsd, tsk_result_ecotox$left, tsk_result_ecotox$right),
drcHelper_tsk = c(tsk_result_drc$LD50, tsk_result_drc$gsd, tsk_result_drc$conf.int[1], tsk_result_drc$conf.int[2]),
Difference = abs(c(tsk_result_ecotox$mu - tsk_result_drc$LD50,
tsk_result_ecotox$gsd - tsk_result_drc$gsd,
tsk_result_ecotox$left - tsk_result_drc$conf.int[1],
tsk_result_ecotox$right - tsk_result_drc$conf.int[2])),
Validated = abs(c(tsk_result_ecotox$mu - tsk_result_drc$LD50,
tsk_result_ecotox$gsd - tsk_result_drc$gsd,
tsk_result_ecotox$left - tsk_result_drc$conf.int[1],
tsk_result_ecotox$right - tsk_result_drc$conf.int[2])) < 0.0001
)
library(kableExtra)
kable(comparison_table) %>%
kable_styling(bootstrap_options = c("striped", "hover")) %>%
row_spec(which(!comparison_table$Validated), background = "#FFCCCC")| Metric | ecotoxicology_TSK | drcHelper_tsk | Difference | Validated |
|---|---|---|---|---|
| LD50/Mu | 43.893393 | 43.893393 | 0 | TRUE |
| GSD | 1.017763 | 1.017763 | 0 | TRUE |
| CI Lower | 42.404509 | 42.404509 | 0 | TRUE |
| CI Upper | 45.434553 | 45.434553 | 0 | TRUE |
Compare with Probit GLM and DRC
# Probit GLM (binomial family)
model_glm_log10 <- glm(cbind(r, n - r) ~ log10(x), family = binomial(link = "probit"), data = data)
#> Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
summary(model_glm_log10)
#>
#> Call:
#> glm(formula = cbind(r, n - r) ~ log10(x), family = binomial(link = "probit"),
#> data = data)
#>
#> Coefficients:
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) -73.32 76403.95 -0.001 0.999
#> log10(x) 46.08 49100.87 0.001 0.999
#>
#> (Dispersion parameter for binomial family taken to be 1)
#>
#> Null deviance: 9.4482e+01 on 4 degrees of freedom
#> Residual deviance: 4.6395e-10 on 3 degrees of freedom
#> AIC: 5.9464
#>
#> Number of Fisher Scoring iterations: 25
lc50_glm_log10 <- dose.p(model_glm_log10, cf = c(1,2), p = 0.5)
print("GLM Probit LC50 (with SE):")
#> [1] "GLM Probit LC50 (with SE):"
print(lc50_glm_log10)
#> Dose SE
#> p = 0.5: 1.591216 37.46047
lc50_glm_log10_val <- 10^lc50_glm_log10[1]
lc50_glm_log10_val
#> p = 0.5:
#> 39.01356
lc50_glm_log10_se <- attr(lc50_glm_log10, "SE")[1]
a_glm_log10 <- coef(model_glm_log10)[1]
b_glm_log10 <- coef(model_glm_log10)[2]
lc50_glm_log10 == -a_glm_log10/b_glm_log10
#> p = 0.5:
#> TRUE
model_glm_log <- glm(cbind(r, n - r) ~ log(x), family = binomial(link = "probit"), data = data)
#> Warning: glm.fit: fitted probabilities numerically 0 or 1 occurred
lc50_glm_log <- dose.p(model_glm_log, cf = c(1, 2), p = 0.5)
lc50_glm_log_val <- exp(lc50_glm_log[1])
lc50_glm_log_se <- attr(lc50_glm_log, "SE")[1]
a_glm_log <- coef(model_glm_log)[1]
b_glm_log <- coef(model_glm_log)[2]
a_glm_log
#> (Intercept)
#> -73.32047
a_glm_log10
#> (Intercept)
#> -73.32047
# Probit DRC (LN.2 with binomial)
model_drc <- drm(r/n ~ x, fct = LN.2(), weight = n, type = "binomial", data = data)
summary(model_drc)
#>
#> Model fitted: Log-normal with lower limit at 0 and upper limit at 1 (2 parms)
#>
#> Parameter estimates:
#>
#> Estimate Std. Error t-value p-value
#> b:(Intercept) 14.168 63.140 0.2244 0.82245
#> e:(Intercept) 40.338 20.573 1.9607 0.04991 *
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
ed50_drc <- ED(model_drc, 50, interval = "inv") ## note. if here using delta, very wide CI.
#>
#> Estimated effective doses
#>
#> Estimate Lower Upper
#> e:1:50 40.338 36.681 48.478
ED(model_drc, 50, interval = "delta")
#>
#> Estimated effective doses
#>
#> Estimate Std. Error Lower Upper
#> e:1:50 40.337797 20.572946 0.015564 80.660031
print("DRC Probit ED50/LC50 (with CI):")
#> [1] "DRC Probit ED50/LC50 (with CI):"
print(ed50_drc)
#> Estimate Lower Upper
#> e:1:50 40.3378 36.68111 48.47803
drc_lc50 <- ed50_drc[1, "Estimate"]
drc_ci <- c(ed50_drc[1, "Lower"], ed50_drc[1, "Upper"])
model_drc_loge <- drm(r/n ~ log(x), fct = LN.2(), weight = n, type = "binomial", data = data)
ED(model_drc_loge, 50, interval = "delta")
#>
#> Estimated effective doses
#>
#> Estimate Std. Error Lower Upper
#> e:1:50 3.69711 0.32517 3.05979 4.33442
ED(model_drc_loge, 50, interval = "fls")
#>
#> Estimated effective doses
#>
#> Estimate Lower Upper
#> e:1:50 40.330 21.323 76.281
library(knitr)
library(kableExtra)
comparison_table <- data.frame(
Method = c("Trimmed Spearman-Karber (TSK)", "Probit GLM (log10)", "Probit GLM (log)", "Probit DRC (LN.2, delta)", "Probit DRC (LN.2, inv)"),
LC50 = c(43.89339, 39.01356, 39.01356, 40.3378,40.3378),
Lower_95_CI = c(42.40451, NA, NA, 0.015564,36.681),
Upper_95_CI = c(45.43455, NA, NA, 80.66003,48.478),
#SE = c(NA, 37.46047, lc50_glm_log_se, NA),
Notes = c("Non-parametric, robust to model fit issues",
"Fitting issues (probabilities 0 or 1 warning)",
"Fitting issues (probabilities 0 or 1 warning)",
"If using delta method, wide CI due to sparse data at high doses",
"robust to sparse data with inverse regression CI")
)
kable(comparison_table, format = "markdown", digits = 3) %>%
kable_styling(bootstrap_options = c("striped", "hover")) %>%
column_spec(5, width = "4.5in")| Method | LC50 | Lower_95_CI | Upper_95_CI | Notes |
|---|---|---|---|---|
| Trimmed Spearman-Karber (TSK) | 43.893 | 42.405 | 45.435 | Non-parametric, robust to model fit issues |
| Probit GLM (log10) | 39.014 | NA | NA | Fitting issues (probabilities 0 or 1 warning) |
| Probit GLM (log) | 39.014 | NA | NA | Fitting issues (probabilities 0 or 1 warning) |
| Probit DRC (LN.2, delta) | 40.338 | 0.016 | 80.660 | If using delta method, wide CI due to sparse data at high doses |
| Probit DRC (LN.2, inv) | 40.338 | 36.681 | 48.478 | robust to sparse data with inverse regression CI |
The TSK and DRC estimates are close (43.89 vs. 40.34), suggesting robustness, while GLM’s estimate (39.01) with a large SE (SE = 37.46047) indicates fitting issues (caused by fitted probabilities numerically 0 or 1 occurred ). Given the warning in GLM, I’d trust TSK or DRC more for this dataset.
What Kind of Data Should Be Used
TSK is designed for quantal (binary) bioassay data from toxicity tests, such as acute lethality studies in aquatic organisms (e.g., fish, Daphnia) or other dose-response experiments. Key characteristics:
-
Data Structure:
- Doses (
x): A vector of increasing positive concentrations (e.g., chemical levels in mg/L). Zero-dose controls are allowed but handled specially. - Responses (
r): Counts of affected subjects (e.g., dead organisms) at each dose. - Sample sizes (
n): Number of subjects tested per dose (can be constant or vary). - Proportions (
p = r/n): Should ideally span near 0 to near 1 across doses, but trimming allows for incomplete coverage if A is adjusted.
- Doses (
-
Requirements:
- At least one dose with partial response (0 < p < 1) for meaningful estimation.
- Doses in strictly increasing order; responses non-negative and ≤ n.
- Suitable for small to moderate sample sizes (e.g., n=10-20 per dose, 4-6 doses).
- Assumes logarithmic spacing for symmetry, so concentrations (not logs) are input.
- Typical Applications: Data from experiments where responses are binomial (e.g., dead/alive) and doses are discrete. It’s robust to irregular dose spacing and doesn’t require normality assumptions.
If responses don’t cover A to 1-A, the code suggests increasing A or warns of invalid trims.
Comparison to Standard Probit Analysis
Here I wil compare the tsk function from
drcHelper with the Probit GLM and Probit DRC approaches for
estimating LC50 (or ED50) values. I’ll also explain the differences and
similarities among these three methods and provide details on
transforming results between Probit GLM and Probit DRC.
Overview of the Three Methods
-
Trimmed Spearman-Karber (TSK) -
drcHelper::tsk:- Method: The Trimmed Spearman-Karber method is a non-parametric approach for estimating the median lethal concentration (LC50) or effective dose (ED50) from dose-response data. It calculates the LC50 as a weighted average of dose levels where the response transitions from below to above 50%, with optional trimming of extreme data points to reduce the influence of outliers.
-
Implementation: In
drcHelper::tsk, the method uses logarithmic doses by default (use.log.doses = TRUE) and implements the pool-adjacent-violators algorithm (PAVA) viaisotone::gpavato smooth non-monotonic responses. It also calculates confidence intervals based on the variance of the lognormal distribution. - Key Features: No assumption of a specific dose-response model; robust to model misspecification; handles non-monotonic data through smoothing.
-
Probit GLM (Generalized Linear Model with Binomial Family):
- Method: This is a parametric approach using a generalized linear model with a binomial family and probit link function. It models the probability of response (e.g., mortality) as a function of the log-dose, assuming the response follows a normal distribution on the probit scale.
-
Implementation:
glm(cbind(r, n - r) ~ log10(x), family = binomial(link = "probit"))fits the model, andMASS::dose.pestimates the LC50. The model assumes a linear relationship between the log-dose and the probit-transformed response probability. - Key Features: Assumes a specific (normal) distribution for the response; provides standard errors and confidence intervals based on asymptotic theory; sensitive to model fit and data distribution.
-
Probit DRC (Dose-Response Curve with
drcPackage, LN.2 Model):-
Method: This is also a parametric approach but uses
the
drcpackage to fit a log-normal dose-response curve (LN.2 model) with a binomial response type. It models the dose-response relationship using a two-parameter log-normal function, where the response is bounded between 0 and 1. -
Implementation:
drm(r/n ~ x, fct = LN.2(), weight = n, type = "binomial")fits the model, andED()estimates the ED50 (LC50). It uses a log-dose internally for fitting. - Key Features: Similar to Probit GLM but specifically tailored for dose-response analysis; assumes a log-normal distribution; provides confidence intervals via the delta method.
-
Method: This is also a parametric approach but uses
the
Similarities Among the Three Methods
- Objective: All three methods aim to estimate the LC50 (or ED50), the dose at which 50% of the population responds (e.g., mortality or effect).
- Data Input: They all work with binomial data (successes and totals, e.g., number of deaths out of total organisms) and dose levels.
-
Dose Transformation: Both Probit GLM and TSK (by
default) use logarithmic transformations of doses (
log10(x)in GLM,log10internally intsk), and DRC uses log-doses internally for fitting. - Confidence Intervals: All methods provide some form of uncertainty estimate (confidence intervals or standard errors) around the LC50 estimate, though the calculation methods differ.
Differences Among the Three Methods
-
Model Assumption:
- TSK: Non-parametric; no assumption about the underlying dose-response curve shape beyond a general monotonic trend (smoothed if necessary).
- Probit GLM: Parametric; assumes a linear relationship on the probit scale, implying a normal distribution of tolerance thresholds.
- Probit DRC (LN.2): Parametric; assumes a log-normal dose-response curve, which may better fit certain biological responses but still imposes a specific functional form.
-
Handling of Data:
- TSK: Uses trimming to reduce the impact of extreme data points and smoothing (PAVA) to handle non-monotonic responses, making it more robust to data irregularities.
- Probit GLM: Sensitive to deviations from normality on the probit scale; can fail to converge or give unreliable estimates if the data doesn’t fit the model (as seen in your warning about fitted probabilities being 0 or 1).
- Probit DRC: More flexible than GLM for dose-response data due to its specific curve-fitting approach but still assumes a log-normal relationship.
-
Estimation and Output:
- TSK: Directly estimates LC50 as a weighted midpoint between doses, with variance-based confidence intervals.
- Probit GLM: Estimates LC50 via inverse prediction from the fitted linear model on the probit scale; standard errors are based on the model’s covariance matrix.
- Probit DRC: Estimates ED50 via the fitted log-normal curve parameters, with confidence intervals using the delta method.
-
Robustness and Applicability:
- TSK: More robust to model misspecification since it doesn’t assume a specific curve; suitable for small datasets or when parametric assumptions are questionable.
- Probit GLM: Less robust; can fail with extreme data (e.g., complete separation as in your warning) and requires larger sample sizes for reliable estimates.
- Probit DRC: Tailored for dose-response, often more robust than GLM for toxicological data but still requires reasonable fit to the log-normal model.
-
Results from Your Data:
-
TSK (
drcHelper::tsk): LC50 = 43.89339, CI = (42.40451, 45.43455) - Probit GLM: LC50 = 39.01356, SE = 37.46047 (very wide due to fitting issues)
-
Probit DRC (
drc::LN.2): LC50 = 40.3378, CI = (0.015564, 80.66003) - Observation: TSK and DRC provide similar LC50 estimates (~40-43), while GLM is slightly lower (~39) with a huge SE, indicating fitting problems (likely due to the warning about numerical probabilities of 0 or 1). DRC’s wide CI suggests uncertainty in the upper bound, possibly due to sparse data at higher doses.
-
TSK (
Transformation Between Probit GLM and Probit DRC
Both Probit GLM and Probit DRC model the dose-response relationship on a transformed scale, but they differ in their functional forms and parameterizations. Here’s how to transform or relate results between them:
1. Probit GLM to Probit DRC
-
Probit GLM Model:
The GLM with a probit link models the probability of response as: \[ \mathrm{probit}(p) = a + b \cdot \log_{10}(\mathrm{dose}) \] where \(p\) is the proportion affected, \(a\) is the intercept, and \(b\) is the slope.The LC50 (dose at which 50% are affected) is found by solving for dose when \(p = 0.5\): \[ \mathrm{probit}(0.5) = 0 = a + b \cdot \log_{10}(\mathrm{LC}_{50}) \] which gives: \[ \log_{10}(\mathrm{LC}_{50}) = -\frac{a}{b} \]
Probit DRC (LN.2):
The DRC modelLN.2()fits a log-normal curve, parameterized as: \[ p = \Phi\left( \frac{\log(\mathrm{dose}) - \log(e)}{1/b} \right) \] where \(e\) is the ED50 (LC50) parameter, \(b\) is the slope, and \(\Phi\) is the standard normal CDF. Here, \(\log\) denotes the natural logarithm (\(\ln\)), not \(\log_{10}\).-
Transformation:
To relate the GLM and DRC parameterizations, convert from base-10 to natural logarithm using \(\log(10) \approx 2.302585\):If GLM gives: \[ \log_{10}(\mathrm{LC}_{50}) = -\frac{a}{b} \] then: \[ \ln(\mathrm{LC}_{50}) = \log_{10}(\mathrm{LC}_{50}) \cdot \log(10) = \left(-\frac{a}{b}\right) \cdot 2.302585 \] which approximates the DRC’s \(\log(e)\) parameter.
The slope parameter transforms as: \[ b_{\mathrm{DRC}} \approx \frac{1}{b_{\mathrm{GLM}} \cdot \log(10)} \] since the DRC slope is inversely related to the spread on the log scale.
b_drc <- coef(model_drc)[1]
-log(coef(model_drc)[2])*b_drc - a_glm_log
#> e:(Intercept)
#> 20.93709
b_glm_log*log(10) - b_glm_log10
#> log(x)
#> -8.05251e-10
b_drc
#> b:(Intercept)
#> 14.16805
b_glm_log
#> log(x)
#> 20.01154
coef(model_drc_loge)[1]
#> b:(Intercept)
#> 51.66112The DRC LN.2 slope parameter is on the natural log scale and should equal to the GLM (log) slope.
-
Practical Note:
Due to different fitting algorithms and assumptions, exact equivalence is not guaranteed. In your data, GLM’s LC50 (39.01356) and DRC’s LC50 (40.3378) are close but not identical, reflecting slight differences in model fit.
2. Probit DRC to Probit GLM
-
Reverse Transformation:
If DRC gives \(e\) (LC50) on the original scale, compute: \[ \ln(e) \] and convert to base-10 log as: \[ \log_{10}(e) = \frac{\ln(e)}{\log(10)} \]
In the example dataset:
-a_glm_log10/b_glm_log10*log(10)
#> (Intercept)
#> 3.663909
-a_glm_log/b_glm_log
#> (Intercept)
#> 3.663909
log(coef(model_drc)[2])
#> e:(Intercept)
#> 3.697289Then, approximate GLM parameters as: \[ b_{\mathrm{GLM}} \approx \frac{1}{b_{\mathrm{DRC}} \cdot \log(10)} \] \[ a_{\mathrm{GLM}} \approx -b_{\mathrm{GLM}} \cdot \log_{10}(e) \]
Again, exact matches are unlikely due to fitting differences.
-
Practical Note:
Use \(\log_{10}(x)\) in GLM to align with DRC’s internal log-scale fitting, though DRC uses \(\ln\) internally.
Recommendations and Conclusion
-
Choosing a Method:
- Use TSK (
drcHelper::tsk) when data is sparse, non-monotonic, or when you want a non-parametric approach robust to model misspecification. It’s ideal for quick, reliable LC50 estimates without strong distributional assumptions. - Use Probit GLM for larger datasets where a normal tolerance distribution is plausible and you need standard statistical inference (e.g., hypothesis testing on coefficients). Be cautious of convergence issues (as in your warning).
- Use Probit DRC (LN.2) for dose-response-specific analysis, especially when a log-normal model fits the biological response better. It’s more tailored to toxicology than GLM but still assumes a specific curve.
- Use TSK (
-
Transformation Caveat: While theoretical
transformations between GLM and DRC are possible, practical differences
in optimization and model assumptions mean direct conversion isn’t
exact. Use the respective package functions (
dose.pfor GLM,EDfor DRC) for accurate estimates.
Example where results agree
The differences in parameter estimates between probit GLM and drc::LN.2 (and even TSK) for the previous example dataset are due to the extreme separation and lack of intermediate responses—most doses have either 0 or 100% response, with only one intermediate point. This leads to instability and poor identifiability in parametric models, which is why we saw huge standard errors and odd parameter values.
Let’s see what happens with a more “ideal” dataset, such as your
finney71 example, which has a smooth, monotonic
increase in response and a good spread of intermediate response
rates:
finney71 <- data.frame(
dose = c(10.2, 7.7, 5.1, 3.8, 2.6, 0.0),
total = c(50, 49, 46, 48, 50, 49),
affected = c(44, 42, 24, 16, 6, 0)
)
finney71$prop <- finney71$affected / finney71$total
# Probit GLM (log10)
glm_log10 <- glm(cbind(affected, total - affected) ~ log10(dose), family = binomial(link = "probit"), data = finney71[finney71$dose > 0, ])
lc50_log10 <- MASS::dose.p(glm_log10, cf = c(1,2), p = 0.5)
cat("GLM log10 LC50:", 10^as.numeric(lc50_log10[1]), "\n")
#> GLM log10 LC50: 4.845492
# Probit GLM (log)
glm_log <- glm(cbind(affected, total - affected) ~ log(dose), family = binomial(link = "probit"), data = finney71[finney71$dose > 0, ])
lc50_log <- MASS::dose.p(glm_log, cf = c(1,2), p = 0.5)
cat("GLM log LC50:", exp(as.numeric(lc50_log[1])), "\n")
#> GLM log LC50: 4.845492
# Probit DRC (LN.2)
library(drc)
drc_mod <- drm(affected/total ~ dose, weights = total, fct = LN.2(), type = "binomial", data = finney71)
ed50_drc <- ED(drc_mod, 50, interval = "delta")
#>
#> Estimated effective doses
#>
#> Estimate Std. Error Lower Upper
#> e:1:50 4.84550 0.24654 4.36229 5.32871
cat("DRC LN.2 LC50:", ed50_drc[1, "Estimate"], "\n")
#> DRC LN.2 LC50: 4.845502
# TSK
finney71 <- finney71[order(finney71$dose),]
tsk_res <- drcHelper::tsk(
x = finney71$dose,
r = finney71$affected,
n = finney71$total,
control = 0,
trim = 0.12, # as suggested by the error
conf.level = 0.95,
use.log.doses = TRUE
)
cat("TSK LC50:", tsk_res$LD50, "\n")
#> TSK LC50: 4.779998
#ecotoxicology::TSK(finney71$dose, finney71$affected, finney71$total, A = 0.12, conf = 0.95)Note that we have to trim here. This is because the TSK method assumes that the response curve is monotonic and that the intermediate responses (not exactly 0 or 1) are most informative for estimating the median effective dose (LC50/EC50).
Results Table
comparison <- data.frame(
Method = c("Probit GLM (log10)", "Probit GLM (log)", "Probit DRC (LN.2)", "TSK (trim = 0.12)"),
LC50 = c(4.845492, 4.845492, 4.845502, 4.779998),
CI_Lower = c(NA, NA, 4.36229, NA),
CI_Upper = c(NA, NA, 5.32871, NA),
Std_Error = c(NA, NA, 0.24654, NA),
Notes = c(
"Identical to log, see below",
"Identical to log10",
"Nearly identical to GLM",
"Robust nonparametric method"
)
)
knitr::kable(comparison, digits = 5)| Method | LC50 | CI_Lower | CI_Upper | Std_Error | Notes |
|---|---|---|---|---|---|
| Probit GLM (log10) | 4.84549 | NA | NA | NA | Identical to log, see below |
| Probit GLM (log) | 4.84549 | NA | NA | NA | Identical to log10 |
| Probit DRC (LN.2) | 4.84550 | 4.36229 | 5.32871 | 0.24654 | Nearly identical to GLM |
| TSK (trim = 0.12) | 4.78000 | NA | NA | NA | Robust nonparametric method |
coef(drc_mod)[1]
#> b:(Intercept)
#> 1.829697
coef(glm_log)[2]
#> log(dose)
#> 1.829768
coef(glm_log10)[2]/log(10)
#> log10(dose)
#> 1.829768Why do they agree here?
- The data has a smooth, monotonic, sigmoidal dose-response with several intermediate response rates (not just 0% or 100%).
- There is no “complete separation” (i.e., not all 0s then all 1s), so the GLM and DRC models can estimate both the slope and intercept reliably.
- The TSK method, being nonparametric, will also yield a similar LC50 because the empirical response curve is monotonic and well-behaved.
Summary:
For well-behaved, monotonic dose-response data with intermediate responses, all methods (parametric and nonparametric) yield nearly identical LC50 estimates. This demonstrates the reliability of these methods when the data meet their assumptions. Differences become pronounced only with sparse, non-monotonic, or extreme datasets.