Backaground
Spearman-Karber method is still in use in regulatory Ecotox endpoints derivation under certain scenarios. A more common approach would be derive a EC50 from interpolation by a smoothing line.
Various Implementations
Here is a comparison between
drcHelper::SpearmanKarber_modified, the original
ecotoxicology::SpearmanKarber, and the
drcHelper::tsk_auto (adaptive “trimmed
Spearman–Karber”).
drcHelper::SpearmanKarber_modified
vs. ecotoxicology::SpearmanKarber
drcHelper::SpearmanKarber_modified is based on a
function written by Sarah and Harold. I provided a fixed and refactored
version. This revision corrected several subtle bugs related to loop
indexing, unsafe handling of log10(0), and hard-coded data sizes, making
the function more robust and reliable for generic data.
In summary, key conceptual differences are:
-
Monotonicity Handling: The original
ecotoxicology::SpearmanKarberuses smoothing, whiledrcHelper::SpearmanKarber_modifieduses a more complex method of pruning and pooling data based on an estimated background mortality rate (c) when control mortality is positive. -
Confidence Intervals:
drcHelper::SpearmanKarber_modifiedincorporates Fieller’s Theorem for more accurate confidence intervals when control mortality is present, a feature the original lacks. -
Flexibility:
drcHelper::SpearmanKarber_modifiedwas designed to handle variable totals per dose (n_i), while the original assumes a constantN.
TSK
The first version of the wrapper function tsk_auto only
auto-trims if tsk() throws a specific trim error. In the
case dose 0 is included in the data, tsk() didn’t error but returned a
degenerate estimate (LD50=0, NaN CI). Turning off logs avoids
that.During the comparison I also wrote a small code patch for
drcHelper:::tsk_auto.data.frame that automatically detects
when a log-transform is requested on data with a zero dose and applies a
minimal trim to prevent errors.
drcHelper:::tsk_auto prioritizes a monotone,
well-behaved response profile by trimming tails rather than smoothing
(like the original) or pruning/pooling by a background-rate threshold
(like drcHelper::SpearmanKarber_modified).
Core estimator (all three are Spearman–Karber variants)
- Base SK trapezoid on log-dose:
- \(m \equiv \sum_{i} \Delta p_i \cdot \frac{\log_{10} x_i + \log_{10} x_{i+1}}{2}\), where \(\Delta p_i = p_{i+1}-p_i\).
- \(\mathrm{LC50} = 10^{m}\).
- Differences are how p is preprocessed (Abbott correction, trimming/pruning, smoothing), and how variance/CI are computed.
How each method preprocesses p (monotonicity and control mortality)
- ecotoxicology::SpearmanKarber (original):
- If proportions aren’t monotone, it smooths them to a non-decreasing sequence, then applies Abbott’s correction.
- Single N assumed; uses that in the variance.
- SpearmanKarber_modified (drcHelper):
- Always does Abbott-like correction from the control (explicit).
- \(p_i^{A} = \frac{p_i - p_0}{1 - p_0}\), with \(p_i = r_i/n_i\).
- If p0=0: add “ghost” endpoints with p=0 and p=1 at extrapolated log-doses; integrate over the full [0,1].
- If p0>0: estimate background mortality c by pooled cumulative
minimum:
- \(c = \min_k \frac{\sum_{i=1}^k r_i}{\sum_{i=1}^k n_i}\), with $n_c $the pooled denominator at the argmin.
- Pool doses with p_i<c into the first group; then rescale the
remaining to [0,1]:
- \(\tilde{p}_1=0,\ \tilde{p}_{m+1}=1,\ \tilde{p}_i=\frac{p_i-c}{1-c}\) for interior points.
- Always does Abbott-like correction from the control (explicit).
- drcHelper::tsk_auto (trimmed SK):
- Applies no smoothing or pooling. It trims symmetric tails (proportion t at each end) to ensure monotone increase and adequate range.
- Over the retained window [L, U], renormalizes:
- \(\tilde{p}_i = \frac{p_i - p_L}{p_U - p_L}\), with \(\tilde{p}_L=0,\ \tilde{p}_U=1\).
- The auto-trim tries the smallest t that allows monotone increase and full [0,1] span (as suggested by tsk’s own diagnostic), capped at max.trim.
Estimator details and equations
- Original (after smoothing and Abbott):
- \(m = \sum_{i=1}^{n-1} \Delta p_i \cdot \frac{\log_{10} x_i + \log_{10} x_{i+1}}{2}\), \(\mathrm{LC50} = 10^m\).
- Modified (your function), control mortality = 0:
- Add ghost endpoints: \(p_0=0,\ p_{n+1}=1\), and extrapolated \(\log_{10}x_0,\ \log_{10}x_{n+1}\).
- \(\mu = \sum_{i=0}^{n} (p_{i+1}-p_i)\cdot \frac{\log_{10}x_i+\log_{10}x_{i+1}}{2}\), \(\mathrm{LC50}=10^{\mu}\).
- Modified (your function), control mortality > 0:
- After pruning and rescaling to \(\tilde
p\), compute raw trapezoid:
- \(\mu^{\mathrm{raw}} = \sum_{i=1}^{m} (\tilde{p}_{i+1}-\tilde{p}_i)\cdot \frac{\log_{10}x_i+\log_{10}x_{i+1}}{2}\).
- Adjust back for c using the first-segment midpoint \(a = \frac{\log_{10}x_0+\log_{10}x_1}{2}\):
- \(\mu = \frac{\mu^{\mathrm{raw}} - c\,a}{1 - c}\), \(\mathrm{LC50}=10^\mu\).
- After pruning and rescaling to \(\tilde
p\), compute raw trapezoid:
- tsk_auto (trimmed SK):
- With indices L..U after trim and renormalized \(\tilde p\):
- \(\mu_{TSK} = \sum_{i=L}^{U-1} (\tilde{p}_{i+1}-\tilde{p}_i)\cdot \frac{\log_{10}x_i+\log_{10}x_{i+1}}{2}\),
- \(\mathrm{LC50}=10^{\mu_{TSK}}\).
- With indices L..U after trim and renormalized \(\tilde p\):
Variance and confidence intervals
- Original
ecotoxicology::SpearmanKarber:- $V_m = _{i=2}^{n-1} p_i(1-p_i) $(constant N).
- 95% CI: \(m \pm 2\sqrt{V_m}\); back-transform to LC50.
- Modified
drcHelper::SpearmanKarber_modified, p0=0:- Per-dose n_i: \(\operatorname{Var}(\mu) = \sum_{i=2}^{n-1} 0.25(\log_{10}x_{i-1}-\log_{10}x_{i+1})^2 \frac{p_i(1-p_i)}{n_i}\).
- CI: \(\mu \pm z_{1-\alpha/2}\sqrt{\operatorname{Var}(\mu)}\).
- Modified
drcHelper::SpearmanKarber_modified, p0>0 (Fieller on log-scale):- Define \(V_{12} = \left(\frac{\log_{10} x_0 + \log_{10} x_1}{2}\right)^2 \frac{c(1-c)}{n_c}\), \(V_{22} = \frac{c(1-c)}{n_c}\), \(V_{11} = \operatorname{Var}(\mu)+V_{12}\).
- With \(z=z_{1-\alpha/2}\), \(B=1-c\), \(g=\frac{z^2V_{22}}{B^2}\):
- \(\sigma = \frac{z}{B} \cdot \frac{\sqrt{V_{11}-2\mu V_{12}+\mu^2V_{22}-g\left(V_{11}-\frac{V_{12}^2}{V_{22}}\right)}}{1-g}\),
- CI: \(\mu - \frac{gV_{12}}{V_{22}} \pm \sigma\); back-transform.
- tsk_auto (
drcHelper::tsk_auto, typical TSK):- Uses the same normal-approx SK variance as your p0=0 branch but on
the trimmed-and-renormalized grid:
- \(\operatorname{Var}(\mu_{TSK}) = \sum_{i=L+1}^{U-1} 0.25(\log_{10}x_{i-1}-\log_{10}x_{i+1})^2 \frac{\tilde p_i(1-\tilde p_i)}{n_i}\),
- CI: \(\mu_{TSK} \pm z_{1-\alpha/2}\sqrt{\operatorname{Var}(\mu_{TSK})}\).
- Fieller is generally not used in trimmed SK; CIs are normal-approx based on the trimmed segment.
- Uses the same normal-approx SK variance as your p0=0 branch but on
the trimmed-and-renormalized grid:
Key contrasts: tsk_auto vs modified vs original
- Handling non-monotonicity:
- tsk_auto: trims tails and renormalizes; data-driven trim chosen automatically.
- Modified: either uses ghost endpoints (p0=0) or prunes doses below a data-driven c and rescales (p0>0).
- Original: smooths to enforce monotonic increase then applies Abbott.
- Control mortality:
- tsk_auto: supplies control dose (control=0 by default). The underlying tsk typically corrects via the control or excludes it when trimming. It does not estimate a background c via cumulative pooling.
- Modified: explicit Abbott correction and, if p0>0, a pooled minimum estimator of background c, plus Fieller CIs.
- Original: Abbott after smoothing (single N).
- Log transformation:
- tsk_auto: can turn log-doses on/off via use.log.doses; practical when x includes a 0 dose (control).
- Modified: always uses log10; explicitly creates ghost endpoints to avoid depending on log10(0).
- Original: uses log10 internally; relies on na.rm=TRUE to avoid log10(0) breaks if present.
- Variance/CI:
- tsk_auto: normal-approx on the trimmed/renormalized grid, per-dose n_i.
- Modified: normal-approx for p0=0; Fieller when p0>0.
- Original: constant-N variance and a fixed “2” multiplier for 95% CI.
Practical guidance
- If you want robust, minimal-tuning monotone behavior with potentially messy tails: tsk_auto is convenient; it chooses a trim that makes the estimation feasible and stable, avoids over-influence of outlying tails, and respects variable n_i.
- If nonzero control mortality is a concern and you want a background-rate-aware estimate with Fieller CIs: your modified function targets that scenario directly.
- If your totals are constant and you prefer smoothing to trimming or pruning: the original function is simplest.
How results will typically differ from tsk_auto
- When control mortality > 0:
-
drcHelper::SpearmanKarber_modifiedestimates c by pooled cumulative minima and applies Fieller CIs; tsk_auto trims instead, then uses normal-approx CIs. Estimates and CIs will differ, especially if early-dose mortalities fluctuate around the background.
-
- When tails are noisy:
- tsk_auto will remove tail influence via trim; your modified (p0=0) keeps full range but adds ghost endpoints; the original smooths instead. Trimming usually reduces variance and sensitivity to extreme tail spacing.
- With variable totals n_i:
- Both tsk_auto and
drcHelper::SpearmanKarber_modifieduse per-dose n_i in variance; theecotoxicolog::SpearmanKarberassumes constant N and may misstate precision when n_i vary.
- Both tsk_auto and
## install.packages("isotone")
## devtools::install_github(repo="brsr/tsk")
library(drcHelper)
## library(tsk)
tsk(c(1, 10, 100, 1000), 20, c(0, 3, 17, 20))
#>
#> Trimmed Spearman-Karber method using 0 percent trim
#>
#> Data was not smoothed
#> Calculation done using the logs of the doses
#> Estimated LD50: 31.62278 GSD of estimate: 1.296928
#> 95 percent confidence interval on LD50:
#> 18.99717 52.63942
data(hamilton)
x<-c(0,0.2,0.3,0.375,0.625,2)
n<-c(30,30,30,30,30,30)
r<-c(0,1,3,16,24,30)
skm <- SpearmanKarber_modified(x,r,n)
skm
#> $log10LC50
#> [1] -0.3468666
#>
#> $varianceOfLog10LC50
#> [1] 0.0009713409
#>
#> $StandardDeviationOfm
#> [1] 0.03116634
#>
#> $confidenceIntervalLog10
#> [1] -0.4079515 -0.2857817
#>
#> $LC50
#> [1] 0.449918
#>
#> $confidenceIntervalLC50
#> [1] 0.3908845 0.5178671
#>
#> $conf.level
#> [1] 0.95
tsk_auto(x,n,r, use.log.doses = FALSE)
#>
#> Trimmed Spearman-Karber method using 0 percent trim
#>
#> Data was not smoothed
#> Calculation done using the raw doses (no log transform)
#> Estimated LD50: 0.5620833 SD of estimate: 0.06153895
#> 95 percent confidence interval on LD50:
#> 0.4414692 0.6826975
tsk_auto(x,n,r, use.log.doses = TRUE)
#> tsk_auto: zero dose with log transform detected; applying auto-trim = 0.0333
#>
#> Trimmed Spearman-Karber method using 3.333333 percent trim
#>
#> Data was not smoothed
#> Calculation done using the logs of the doses
#> Estimated LD50: 0.4438306 GSD of estimate: 1.079123
#> 95 percent confidence interval on LD50:
#> 0.3822956 0.5152705
drcHelper::tsk(x, n, r, trim = 1/30, use.log.doses = FALSE)
#>
#> Trimmed Spearman-Karber method using 3.333333 percent trim
#>
#> Data was not smoothed
#> Calculation done using the raw doses (no log transform)
#> Estimated LD50: 0.5313244 SD of estimate: 0.06430354
#> 95 percent confidence interval on LD50:
#> 0.4052918 0.6573570
drcHelper::tsk(x, n, r, trim = 1/30, use.log.doses = TRUE)
#>
#> Trimmed Spearman-Karber method using 3.333333 percent trim
#>
#> Data was not smoothed
#> Calculation done using the logs of the doses
#> Estimated LD50: 0.4438306 GSD of estimate: 1.079123
#> 95 percent confidence interval on LD50:
#> 0.3822956 0.5152705
## ecotoxicology::TSK errored for A=0, suggesting A ≥ 1/30 ≈ 0.033333… to make the response span usable for trimming.
## A = 0.0333 failed due to a formatting bug in that version; use A that makes A*100 an integer (e.g., 0.04).
ecotoxicology::TSK(x*100, r, n, A = 0.04, conf = 0.95)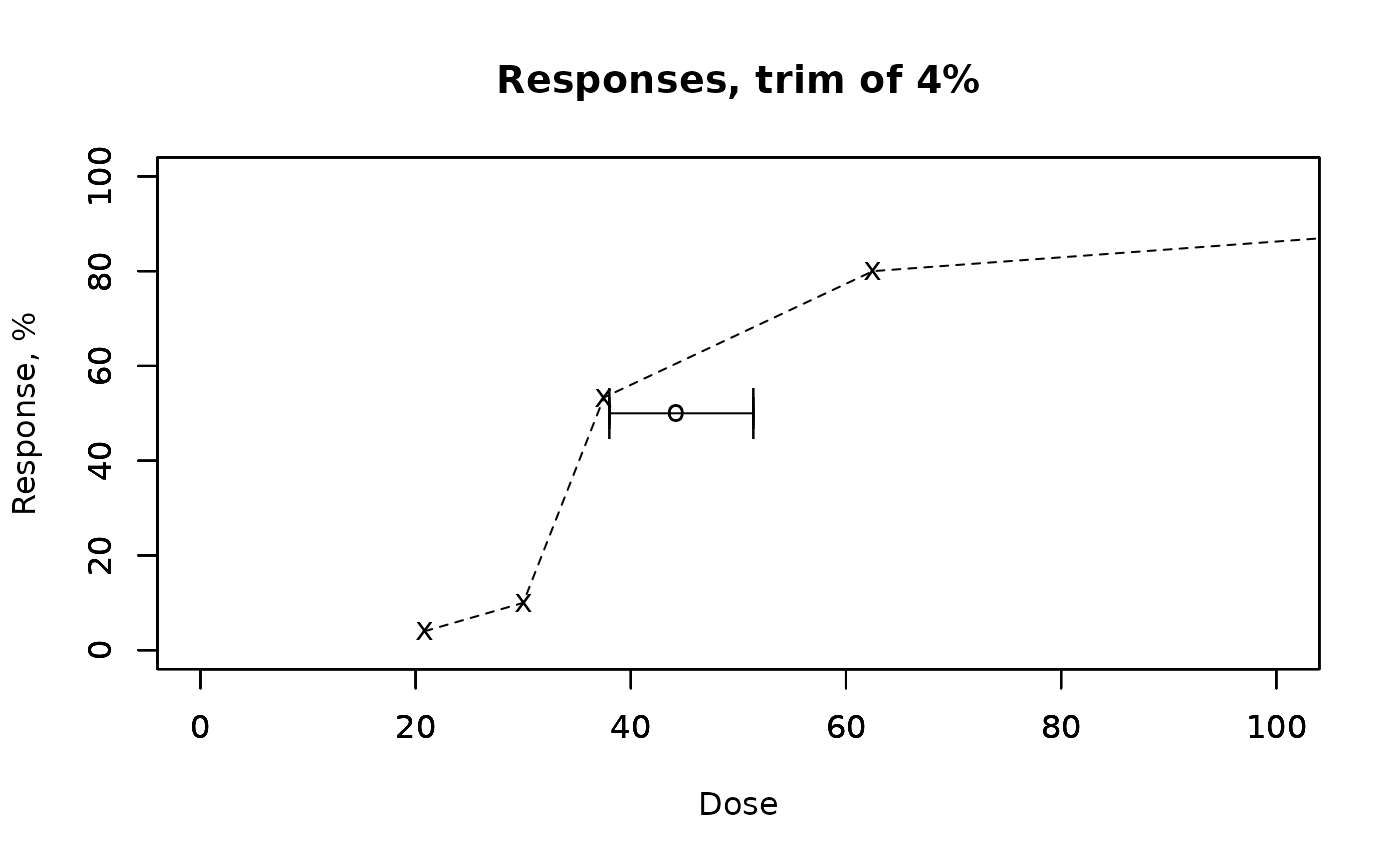
#> $mu
#> [1] 44.21762
#>
#> $gsd
#> x
#> 1.079643
#>
#> $left
#> x
#> 38.05109
#>
#> $right
#> x
#> 51.3835
p <- r/n
x2 <- x
x2[1] <- 1e-1 ## replace it with very small values.
ecotoxicology::SpearmanKarber(cbind(x2,r,p),N=30, retData = FALSE, showOutput = TRUE,showPlot = FALSE)
#> x2 r p smoothedAdjusted
#> [1,] 0.100 0 0.00000000 0.00000000
#> [2,] 0.200 1 0.03333333 0.03333333
#> [3,] 0.300 3 0.10000000 0.10000000
#> [4,] 0.375 16 0.53333333 0.53333333
#> [5,] 0.625 24 0.80000000 0.80000000
#> [6,] 2.000 30 1.00000000 1.00000000
#> log10 LC50 = -0.3489489
#> estimated variance of m = 0.001146069
#> 95% confidence interval for m = -0.416656192612695 -0.281241619158308
#> estimated LC50 = 0.447766
#> estimated 95% confidence interval for the estimated LC50 = 0.383127924893929 0.523309213196528
ecotoxicology::SpearmanKarber(cbind(x2,r,n),N=30, retData = FALSE, showOutput = TRUE,showPlot = FALSE)
#> x2 r n smoothedAdjusted
#> [1,] 0.100 0 30 0
#> [2,] 0.200 1 30 0
#> [3,] 0.300 3 30 0
#> [4,] 0.375 16 30 0
#> [5,] 0.625 24 30 0
#> [6,] 2.000 30 30 0
#> log10 LC50 = 0
#> estimated variance of m = 0
#> 95% confidence interval for m = 0 0
#> estimated LC50 = 1
#> estimated 95% confidence interval for the estimated LC50 = 1 1
## remove 0 dose
ecotoxicology::SpearmanKarber(cbind(x[-1],r[-1],p[-1]),N=30, retData = TRUE, showOutput = TRUE,showPlot = FALSE)
#> smoothedAdjusted
#> [1,] 0.200 1 0.03333333 0.00000000
#> [2,] 0.300 3 0.10000000 0.06896552
#> [3,] 0.375 16 0.53333333 0.51724138
#> [4,] 0.625 24 0.80000000 0.79310345
#> [5,] 2.000 30 1.00000000 1.00000000
#> log10 LC50 = -0.331689
#> estimated variance of m = 0.001126808
#> 95% confidence interval for m = -0.398824952238237 -0.264553128754662
#> estimated LC50 = 0.4659196
#> estimated 95% confidence interval for the estimated LC50 = 0.399185766700702 0.543809601530862
#> $log10LC50
#> [1] -0.331689
#>
#> $varianceOfm
#> [1] 0.001126808
#>
#> $confidenceInterval95m
#> [1] -0.3988250 -0.2645531
#>
#> $LC50
#> [1] 0.4659196
#>
#> $confidenceInterval95LC50
#> [1] 0.3991858 0.5438096
x=c(0,15.54,20.47,27.92,35.98,55.52)
n=c(40,40,40,40,40,40)
r=c(3,2,6,11,18,33)
ecotoxicology::SpearmanKarber(cbind(x,r,n),N=30, retData = FALSE, showOutput = TRUE,showPlot = FALSE)
#> x r n smoothed smoothedAdjusted
#> [1,] 0.00 3 40 0.08333333 0.0000000
#> [2,] 15.54 2 40 0.08333333 0.0000000
#> [3,] 20.47 6 40 0.20000000 0.1272727
#> [4,] 27.92 11 40 0.36666667 0.3090909
#> [5,] 35.98 18 40 0.60000000 0.5636364
#> [6,] 55.52 33 40 1.10000000 1.1090909
#> log10 LC50 = 1.692103
#> estimated variance of m = 0.0002847302
#> 95% confidence interval for m = 1.65835487286972 1.72585067406059
#> estimated LC50 = 49.2156
#> estimated 95% confidence interval for the estimated LC50 = 45.5359994197084 53.1925332910634
# Your data
x <- c(0, 0.2, 0.3, 0.375, 0.625, 2)
n <- c(30, 30, 30, 30, 30, 30)
r <- c(0, 1, 3, 16, 24, 30)
# Create a master comparison function
tidy_compare <- function(x, n, r, conf.level = 0.95) {
# A list to hold the output rows
all_rows <- list()
# Call each extract function and add the result to the list
all_rows[[length(all_rows) + 1]] <- drcHelper:::extract_SpearmanKarber_modified(x, n, r, conf.level)
all_rows[[length(all_rows) + 1]] <- drcHelper:::extract_tsk_auto(x, n, r, use.log.doses = TRUE, conf.level)
all_rows[[length(all_rows) + 1]] <- drcHelper:::extract_tsk_auto(x, n, r, use.log.doses = FALSE, conf.level)
all_rows[[length(all_rows) + 1]] <- drcHelper:::extract_ecotox_SpearmanKarber(x, n, r)
# You can easily add more functions here, e.g.:
# all_rows[[length(all_rows) + 1]] <- extract_ecotox_TSK(x, n, r, conf = conf.level)
# Combine all the single-row data frames into one table
do.call(rbind, all_rows)
}
# Run the comparison
comparison_table <- tidy_compare(x, n, r)
#> tsk_auto: zero dose with log transform detected; applying auto-trim = 0.0333
# Print the final result
## print(comparison_table)
comparison_table %>% knitr::kable(.,digits = 3)| method | scale | trim_or_A | LC50 | LCL | UCL | log10LC50 | SD_LC50 | SD_log10 | GSD | notes |
|---|---|---|---|---|---|---|---|---|---|---|
| SpearmanKarber_modified | log-dose | NA | 0.450 | 0.391 | 0.518 | -0.347 | NA | 0.031 | NA | |
| tsk_auto | log-dose | 0.033 | 0.444 | 0.382 | 0.515 | -0.353 | NA | 0.076 | 1.079 | |
| tsk_auto | linear-dose | 0.000 | 0.562 | 0.441 | 0.683 | -0.250 | 0.062 | NA | NA | |
| ecotox::SpearmanKarber | log-dose | NA | 0.466 | 0.399 | 0.544 | -0.332 | NA | 0.034 | NA |
Using the wrapper analyze_SK function
# Your data
x <- c(0, 0.2, 0.3, 0.375, 0.625, 2)
n <- c(30, 30, 30, 30, 30, 30)
r <- c(0, 1, 3, 16, 24, 30)
# Define the methods you want to run
methods_to_run <- c("SK_modified", "tsk_auto_log", "tsk_auto_linear", "ecotox_SK")
# Use lapply to run the analysis for each method
all_results <- lapply(methods_to_run, function(m) {
# The ... argument lets you pass method-specific options here if needed
# e.g., analyze_SK(x, n, r, method = m, max.trim = 0.4)
analyze_SK(x, n, r, method = m)
})
#> tsk_auto: zero dose with log transform detected; applying auto-trim = 0.0333
# Combine the list of single-row data frames into one final table
comparison_table <- do.call(rbind, all_results)
# Print the result
print(comparison_table)
#> method scale trim_or_A LC50 LCL UCL
#> 1 SK_modified log-dose NA 0.4499180 0.3908845 0.5178671
#> 2 tsk_auto_log log-dose 0.03333333 0.4438306 0.3822956 0.5152705
#> 3 tsk_auto_linear linear-dose 0.00000000 0.5620833 0.4414692 0.6826975
#> 4 ecotox_SK log-dose NA 0.4659196 0.3991858 0.5438096
#> log10LC50 SD_LC50 SD_log10 GSD notes
#> 1 -0.3468666 NA 0.03116634 NA
#> 2 -0.3527827 NA 0.07614879 1.079123
#> 3 -0.2501993 0.06153895 NA NA
#> 4 -0.3316890 NA 0.03356796 NA- All three valid SK variants (your modified, TSK with A ≈ 0.0333 and use.log.doses = FALSE, and original SpearmanKarber with N = 30) should give very similar LC50 near 0.45 because responses increase cleanly from 0 to 1 across doses.
- Minor differences can arise from:
- Whether ghost endpoints or trimming are used,
- Variance/CI formulas (Fieller vs normal approximation),
- Whether logs are used with a zero-dose control.
Notes on GSD and SD_log10
This is related to how we describe uncertainty for log-normal data, and it’s a frequent point of confusion.
In short: They both measure the exact same amount of uncertainty, but they express it on different scales.
-
SD_log10is additive on the log10 scale. -
GSDis multiplicative on the original concentration scale.
SD_log10: The Standard Deviation of the log10(LC50)
This is the more straightforward statistical measure. When we perform Spearman-Karber analysis, we are actually estimating the mean of the distribution of the logarithms of the toxic thresholds.
-
What it is: The standard deviation of our
log10LC50estimate. If our estimate is \(m = \log_{10}(\text{LC50})\), thenSD_log10is the standard deviation of \(m\). - How it’s used: It is used to construct a confidence interval on the log10 scale. You add and subtract this value (multiplied by a z-score like 1.96) to get your confidence limits. \(\text{95% CI for } \log_{10}(\text{LC50}) = m \pm 1.96 \times \text{SD}_{\log_{10}}\)
GSD: The Geometric Standard Deviation
The GSD is a more intuitive way to express this same uncertainty back on the original concentration scale. Since the underlying distribution is log-normal, the spread is multiplicative, not additive.
- What it is: A unitless factor that describes how spread out the data are on the original scale.
-
How it’s derived: It is the exponentiated standard
deviation from the log scale. The relationship depends on the base of
the logarithm used.
- If using natural log (
ln), thenGSD= \(e^{\text{SD}_{\ln}}\). ThedrcHelperpackage uses this convention. - To be consistent with our
SD_log10, we can define a base-10 GSD as: \(\text{GSD} = 10^{\text{SD}_{\log_{10}}}\) And conversely: \(\text{SD}_{\log_{10}} = \log_{10}(\text{GSD})\)
- If using natural log (
- How it’s used: It is used to construct a confidence interval by multiplying and dividing the LC50 estimate. \(\text{95% CI for LC50} = \text{LC50} \times / \div (\text{GSD})^{1.96}\) Which means the lower bound is $ / ()^{1.96} $and the upper bound is \(\text{LC50} \times (\text{GSD})^{1.96}\).
- Analogy: Instead of saying “the true value is plus or minus 10 mg/L,” you’d say “the true value is likely within a factor of 2 of our estimate.” In this case, the GSD would be 2.
Summary Table
| Feature | SD_log10 |
GSD (Geometric Standard Deviation) |
|---|---|---|
| Scale | Logarithmic | Original (Concentration) |
| Operation | Additive (±) | Multiplicative (×/÷) |
| Units | log10(concentration units) | None (it’s a factor) |
| CI on Log Scale | log(LC50) ± z * SD_log10 |
Not used directly |
| CI on Original Scale | Back-transform the log CI | LC50 ×/÷ (GSD)^z |
| Interpretation | “The uncertainty is ±0.03 units on the log10 scale.” | “The uncertainty is by a factor of 1.08 on the original scale.” |
The reason both columns exist in our summary table is that different
functions naturally report one or the other.
SpearmanKarber_modified calculates SD_log10,
while drcHelper::tsk_auto reports the GSD. By
calculating the corresponding value for each, our table allows for a
direct, apples-to-apples comparison of the uncertainty calculated by
each method.
Some Notes
tsk function comes from the R package. Since it is a
single function without dependencies, it is bundled into this helper
package for the purpose of validation and verification.
The original goal of this R package was to replicate the results of a DOS program that used to be provided by the EPA to perform the trimmed Spearman-Karber method. The list “expected” contains the results of that DOS program run on the data from Hamilton et al. Some of these are NA because the EPA program didn’t return a result. The EPA program uses a confidence of 2*pnorm(2)-1=0.9544997 (that is, exactly two sigmas on both sides).
Other Methods
Two commonly used methods for calculating 50% endpoint using serial dilutions are Spearman-Karber method and Reed and Muench method.
The original paper written by Kärber was published in 1931 and is in German [Kärber, G. Archiv f. experiment. Pathol. u. Pharmakol (1931) 162: 480-483 doi:10.1007/BF01863914].
Reed and Muench method
log10 50% end point dilution = log10 of dilution showing a mortality next above 50% - (difference of logarithms × logarithm of dilution factor).
Generally, the following formula is used to calculate “difference of logarithms” (difference of logarithms is also known as “proportionate distance” or “interpolated value”): Difference of logarithms = [(mortality at dilution next above 50%)-50%]/[(mortality next above 50%)-(mortality next below 50%)].
Spearman-Karber method
log10 50% end point dilution = \(- (x_0 - d/2 + d \sum r_i/n_i)\)
\(x_0\) = log10 of the reciprocal of the highest dilution (lowest concentration) at which all animals are positive;
\(d\) = log10 of the dilution factor;
\(n_i\) = number of animals used in each individual dilution (after discounting accidental deaths);
\(r_i\) = number of positive animals (out of \(n_i\)).
Summation is started at dilution \(x_0\).
Newly proposed method Formula 1:
log10 50% end point dilution = -[(total number of animals died/number of animals inoculated per dilution) + 0.5] × log dilution factor.
Formula 2 (if any accidental death occurred):
log10 50% end point dilution = -(total death score + 0.5) × log dilution factor.
References
https://www.cureffi.org/2015/09/20/the-math-behind-spearman-karber-analysis/
Hamilton, M. A.; Russo, R. C.; Thurston, R. V. Trimmed Spearman-Karber Method for Estimating Median Lethal Concentrations in Toxicity Bioassays. Enviro. Sci. Tech. 1977, 11 (7), 714-719. http://dx.doi.org/10.1021/es60130a004
Ibid, 1978, 12 (4), 417. http://dx.doi.org/10.1021/es60140a017